Prostate Cancer
Prostate cancer (PCa) is the most frequently diagnosed cancer among men, with approximately 1.5 million newly diagnosed cases worldwide1. However, although this malignancy is diagnosed in about 1 out of 5 men during their lifetime, 78% survive PCa for 10 years and more.
Diagnosis of PCa is confirmed based on the histopathological verification of tumour presence in prostate biopsies, after a positive result of digital rectal examination (DRE), or after elevated PSA levels and more recently, after a positive or suspicious result of multiparametric magnetic resonance (mpMRI)2.
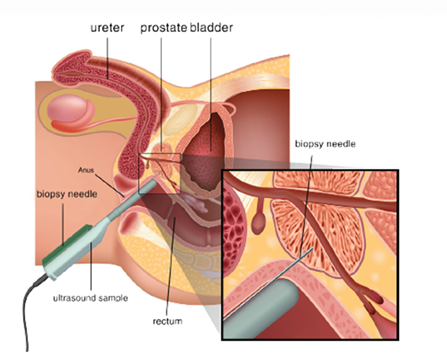
Figure 1. The diagnosis of prostate cancer is based on the invasive biopsies.
Multiple factors not related to prostate malignancy may affect the level of blood PSA (inflammation, infection, presence of benign prostate hyperplasia /BPH etc.). Thus, PSA testing has proven not accurate enough (as specificity is very low). That means that there many false positive results with only less than half (~40%) of all patients with elevated PSA serum levels (≥4 ng/mL) ending up positively confirmed with PCa after biopsy3.
At the same time, introduction of PSA testing and multi-core biopsy resulted in increased detection of slow growing PCa unlikely to progress in the absence of treatment (low risk disease). Uncertainty to properly assess PCa resulted in patients’ overtreatment. This imposes a significant social and economic burden on patients and health-care holders, as a result of the side effects and the high associated treatment costs.

Figure 2. Slow growing PCa is unlikely to progress in the absence of treatment (low risk disease). Selection of patients for active surveillance helps to avoid over-treatment.
Active surveillance is currently a valid alternative to immediate therapy and overtreatment of patients with low risk disease. Active surveillance implies intervention only in patients with local tumour progression.
Better guidance of invasive biopsies through non-invasive means is thus necessary to reduce over-diagnosis and over-treatment4,5,6,7 and guide patients during active surveillance.
Urinary Proteomics Analysis: The CE-MS based PCU Test
Mosaiques applies urinary profiling using capillary electrophoresis online coupled to mass spectrometry (CE-MS) to the non-invasive diagnosis of prostate cancer, to reduce the unnecessary biopsies and patients’ over-treatment and the associated costs. The urinary proteomic profiling of patients with prostate cancer and benign conditions (benign prostate hyperplasia- BPH) enables the identification of specific proteomic biomarkers that are combined in a diagnostic test (PCU -Prostate Check-Up- test) to enable efficient prostate cancer detection.
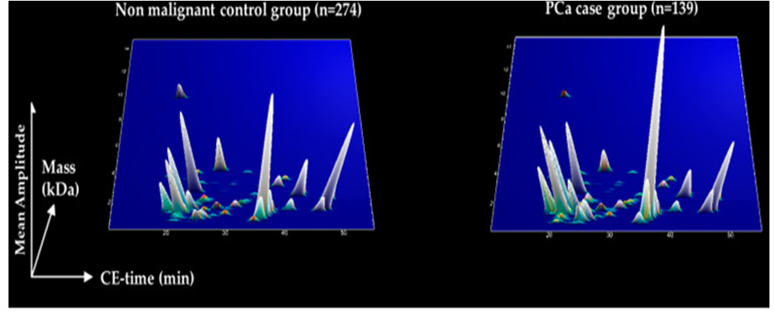
Figure 3. 3D plot of the specific peptide signatures in PCa patients (right panel) and the same signature in male without PCa.
The PCU test simultaneously determines 181 peptide biomarkers indicative for PCa8. The diagnostic accuracy of the PCa proteomic test is developed and validated in a total of over 900 patients8. The sensitivity levels of PCU test are at 93% (87–96; 95% CI;) and the specificity at 69% (63–74; 95% CI), while negative predictive value (NPV) has been estimated at 93.7%8.
Recommendation for patients
The PCU test is recommended for patients with suspicious PSA and/or digital rectal examination results prior to intended biopsy. The test allows distinguishing patients with a positive PSA test who have high probability for PCa from those who do not have it. A positive test result gives a clearer indication for a biopsy than PSA testing alone; whereas due to the high negative predictive value of >90%, in patients with negative test result biopsy is not indicated.
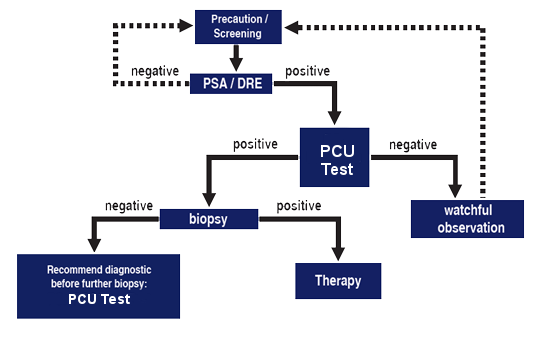
Figure 4. The PCU Test is recommended to correct false positive PSA results.
Benefits of using the PCU Test
The high diagnostic accuracy of PCU test allows to correct the false positive results of PSA testing and continuously can help to reduce the number of painful and unnecessary biopsies as well as related side effects (e.g. bleeding, pain, problems with urination). In addition, severe anxiety in men and their families associated with the false-positive results of the PSA test is an uncomfortable medical condition and can be minimized with the application of the test.
PCU Test is non-invasive and there is no risk associated with sample collection. The benefits associated with the use of this test are further supported by a decision curve analysis demonstrating a high net benefit for biopsy reduction based on the application of the PCU test particularly in the lower range of risk thresholds (<50%), compared to clinical variables such as PSA, PSA density and the European Randomised Study of Screening for Prostate Cancer (ERSPC)8.
Urinary Proteomics Analysis: The CE-MS based PSM Test
About 45% of patients initially diagnosed with prostate cancer have only a slow-growing form of the cancer. This is where active surveillance will be used that only patients with aggressive prostate cancer can be treated. With the protexam PSM test, aggressive prostate cancer can be detected at an early stage.
The urinary proteomic profiling of patients with indolent prostate cancer and those with aggressive forms, enables the identification of specific proteomic biomarkers that are combined in a diagnostic test (PSM-Prostate Status Monitoring test) to enable efficient detection of aggressive prostate cancer.
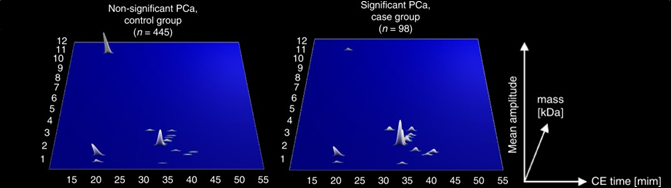
Figure 5. 3D plot of the specific peptide signatures in patients with indolent (non-significant) PCa (left panel) and the same signature in male with aggressive (significant) PCa.
The PSM test simultaneously determines 19 peptide biomarkers indicative for PCa9-10. The diagnostic accuracy of the PSM protexam test is developed and validated in a total of over 900 patients10. The sensitivity levels of PSM test are at 90% and negative predictive value (NPV) has been estimated at 91.8%10.
Recommendation for patients
The PSM test is recommended for patients initially detected with PCa to distinguish aggressive PCa and those that are under AS in order to monitor progression.
Benefits of using the PCU Test
The high diagnostic accuracy of PSM test allows to accurately predict aggressive PCa and continuously can help to reduce the number of painful and unnecessary biopsies as well as related side effects (e.g. bleeding, pain, problems with urination).
REFERENCES
- Sung H., et al. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA A Cancer J. Clin. 2021;71:209–249.
- Mottet N., et al. EAU-EANM-ESTRO-ESUR-SIOG Guidelines on Prostate Cancer-2020 Update. Part 1: Screening, Diagnosis, and Local Treatment with Curative Intent. Eur. Urol. 2021;79:243–262.
- Guichard G., et al. Extended 21-sample needle biopsy protocol for diagnosis of prostate cancer in 1000 consecutive patients. Eur. Urol. 2007;52:430–435.
- Roobol M.J., et al. Screening for prostate cancer: Results of the Rotterdam section of the European randomized study of screening for prostate cancer. Eur. Urol. 2013;64:530–539.
- Arnold M., et al. Recent trends in incidence of five common cancers in 26 European countries since 1988: Analysis of the European Cancer Observatory. Eur. J. Cancer. 2015;51:1164–1187.
- Center M.M., et al. International variation in prostate cancer incidence and mortality rates. Eur. Urol. 2012;61:1079–1092.
- Haas G.P., et al. The worldwide epidemiology of prostate cancer: Perspectives from autopsy studies. Can. J. Urol. 2008;15:3866–3871.
- Frantzi M., et al. Mass Spectrometry-Based Biomarkers to Detect Prostate Cancer: A Multicentric Study Based on Non-Invasive Urine Collection without Prior Digital Rectal Examination. Cancers. 2023 Feb 11;15(4):1166.
- CE-MS-based urinary biomarkers to distinguish non-significant from significant prostate cancer. Frantzi M, Gomez Gomez E, Blanca Pedregosa A, Valero Rosa J, Latosinska A, Culig Z, Merseburger AS, Luque RM, Requena Tapia MJ, Mischak H, Carrasco Valiente J. Br J Cancer. 2019 Jun;120(12):1120-1128.
- Validation of diagnostic nomograms based on CE-MS urinary biomarkers to detect clinically significant prostate cancer. Frantzi M, Heidegger I, Roesch MC, Gomez-Gomez E, Steiner E, Vlahou A, Mullen W, Guler I, Merseburger AS, Mischak H, Culig Z. World J Urol. 2022 Sep;40(9):2195-2203.



